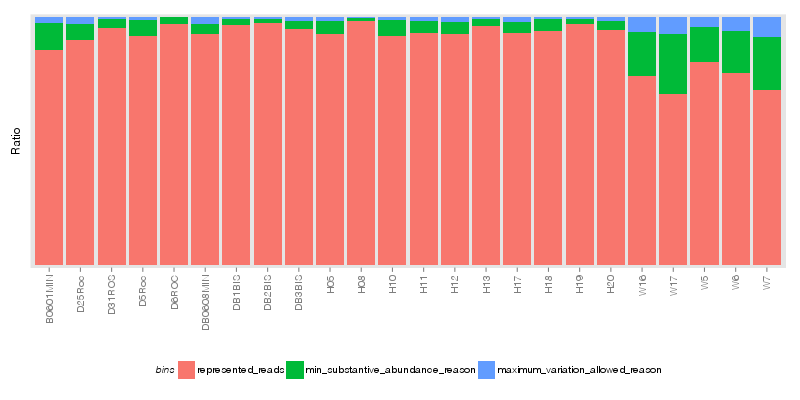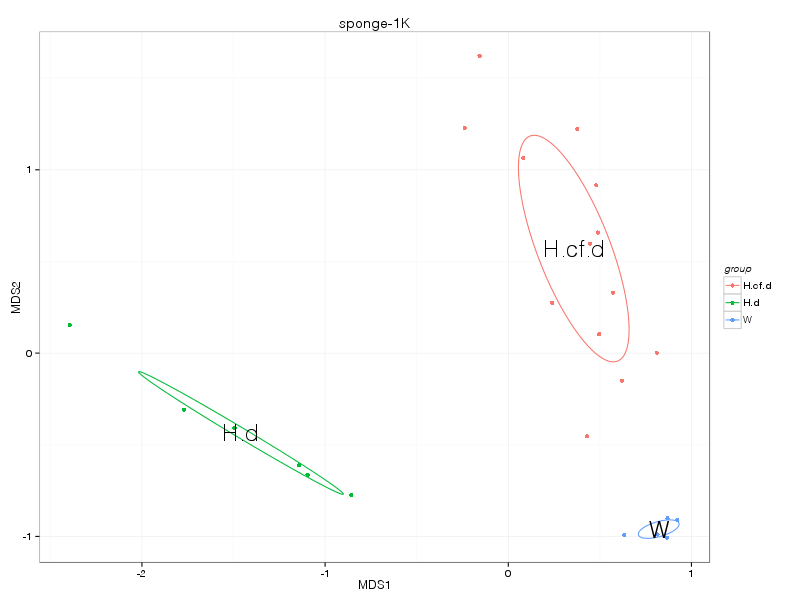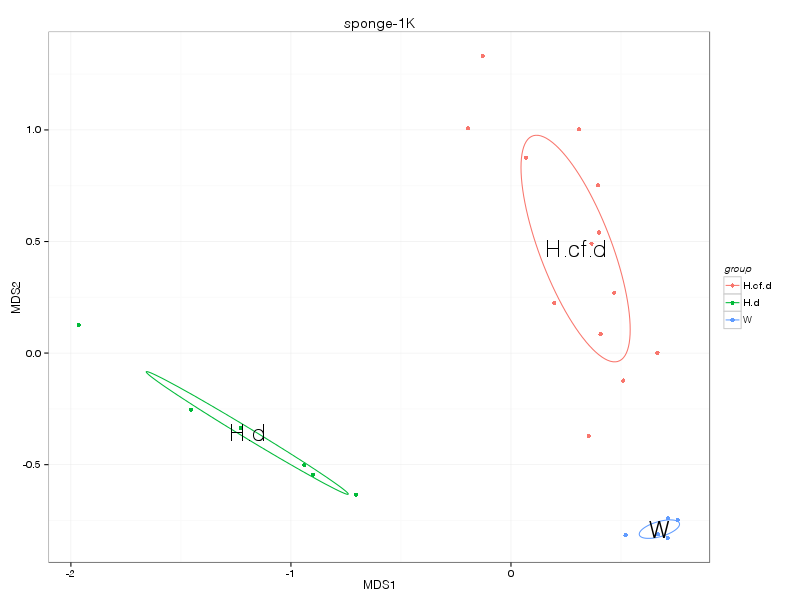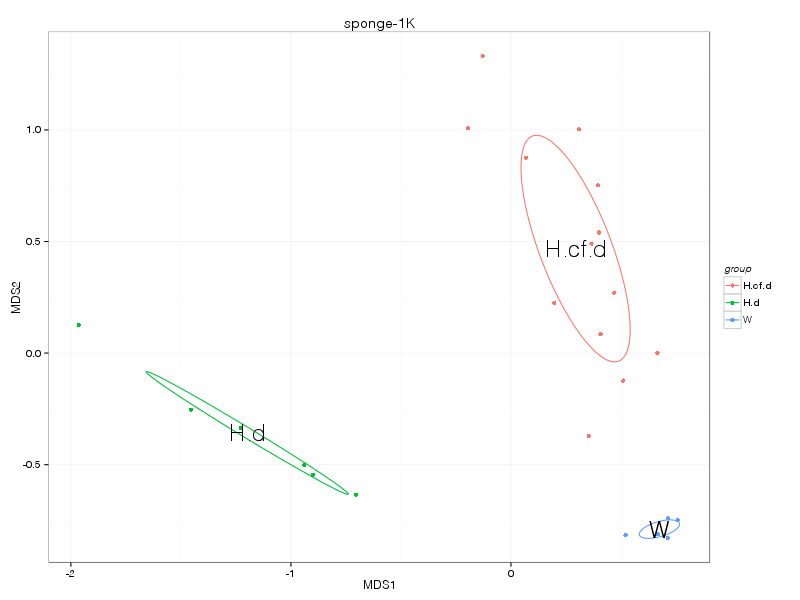
» A summary of what happened.
Minimum Entropy Decomposition analysis was performed on 24,000 read from 24 samples for "sponge-1K" with MED pipeline version 0.1-alpha (available from http://oligotyping.org/MED).
» Meta
| Library version | 0.1-alpha |
| Run date | 03 Nov 14 23:25:00 |
| End of run | 03 Nov 14 23:26:36 |
» Given Parameters
| Min entropy for a component to be picked for decomposition | 0.0965 |
| Perform entropy normalization heuristics | True |
| Max number of discriminants to use for decomposition | 4 |
| Min total abundance of oligotype in all samples | 0 |
| Min substantive abundance of an oligotype (-M) | 5.0 |
| Maximum variation allowed in each node (-V) | 4 nt |
| Nodes agglomerated based on co-occurence patterns | |
| Merge homopolymer splits | False |
| Skip removing outliers | False |
| Try to relocate outliers | False |
» Input Data
| Number of sequences analyzed | 24,000 |
| Number of samples found | 24 |
| Number of characters in each alignment | 402 |
| Average read length (without gaps) | 374 |
» Handling Outliers
| Outliers removed due to -M | 1,759 |
| Outliers removed due to -V | 583 |
| Total number of outliers removed during the refinement | 2,342 |
| Relocated outliers originally removed due to -M | |
| Relocated outliers originally removed due to -V | |
| Total number of relocated outliers | |
| Final number of outliers due to -M | 1,759 |
| Final number of outliers due to -V | 583 |
| Final total number of outliers | 2,342 |
» Nodes
| Number of sequences analyzed | 24,000 |
| Number of sequences represented after quality filtering | 21,658 |
| Number of raw nodes (before the refinement) | 287 |
| Number of final nodes (after the refinement) | 287 |
» Files to analyze results further via third partry applications
| Representative sequences per node | node-representatives.fa.txt |
| Read distribution among samples table | read_distribution.txt |
| Sample/oligotype abundance data matrix (percents) | matrix_percents.txt |
| Sample/oligotype abundance data matrix (counts) | matrix_counts.txt |
| Environment file | environment.txt |
| Mapping file | sample_mapping.txt |
| GEXF file for network analysis | network.gexf |
| Basic topology of MED nodes | topology.gexf |
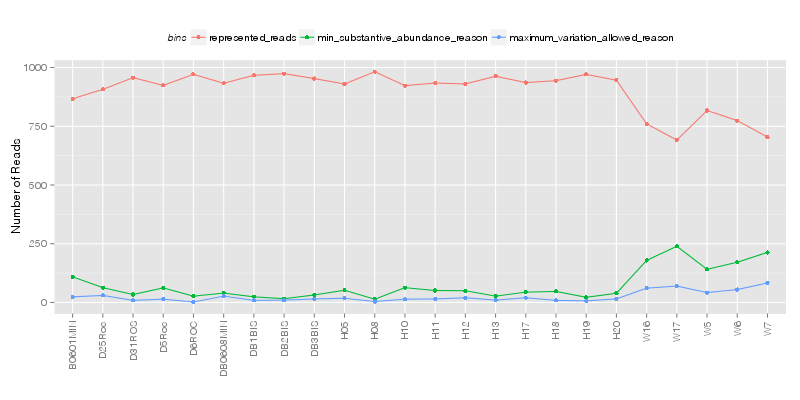
» Total number of reads for each sample that were analyzed.
» Deafult
» cluster_analysis
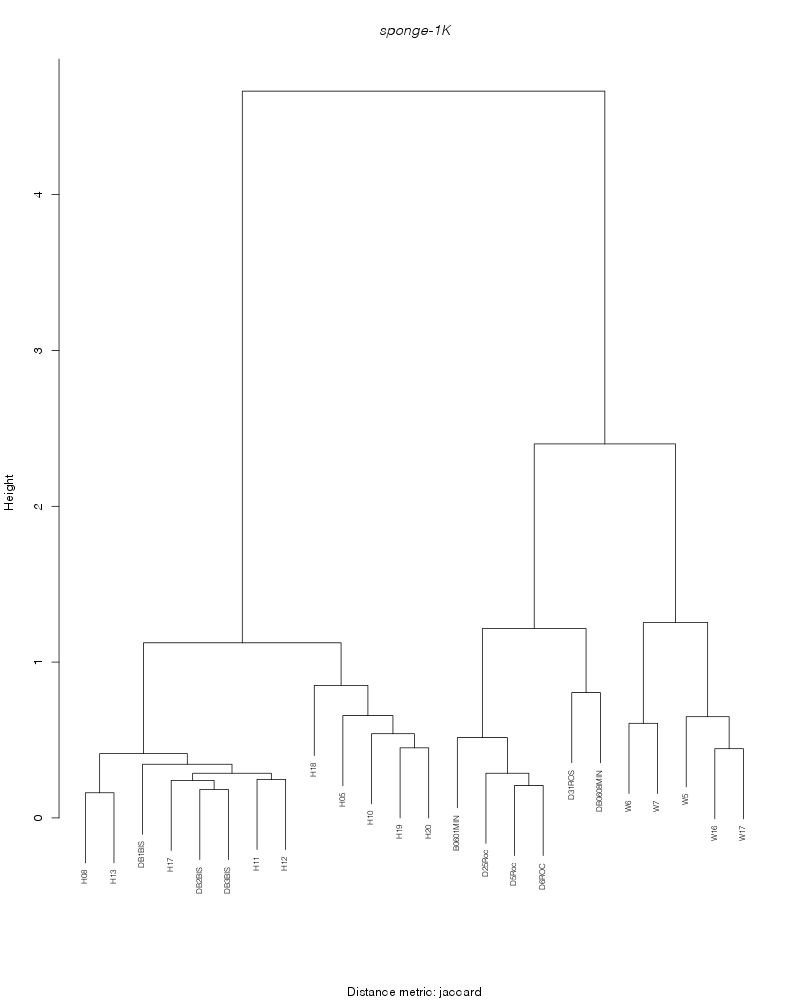
jaccard |
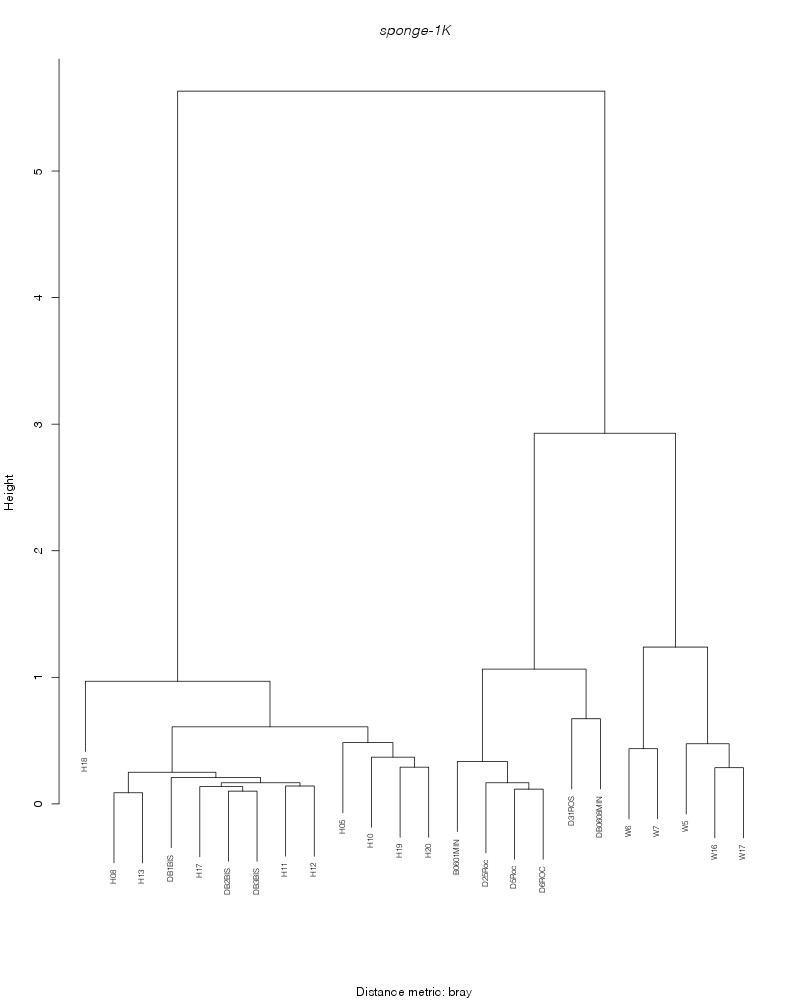
bray |
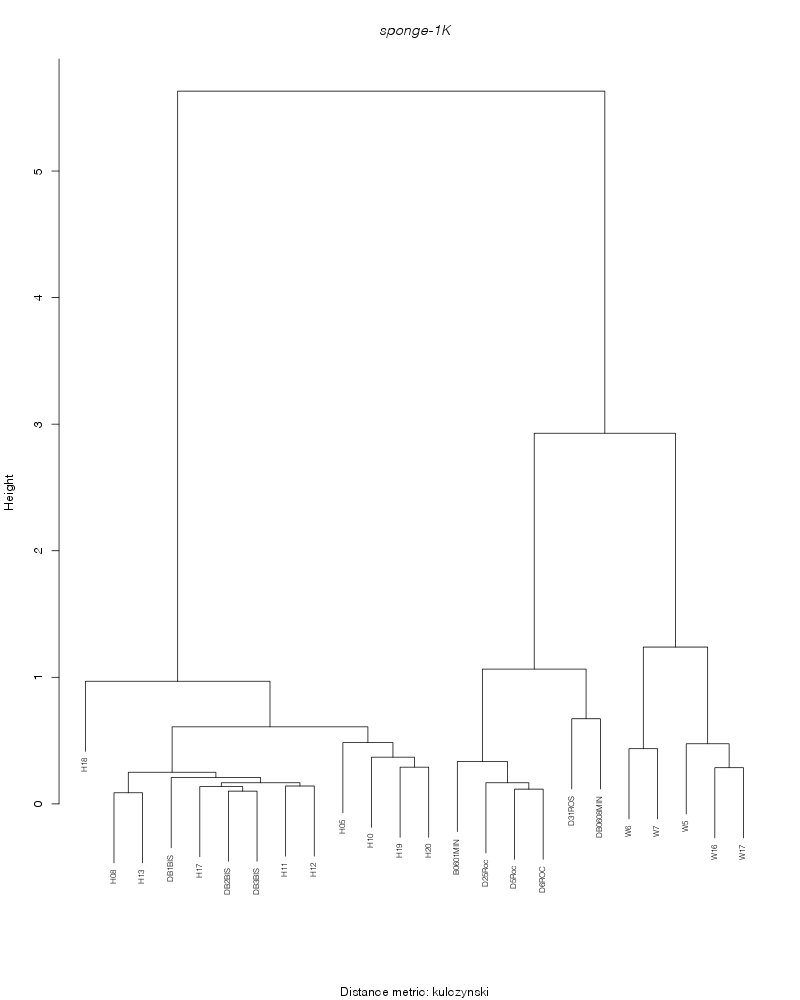
kulczynski |
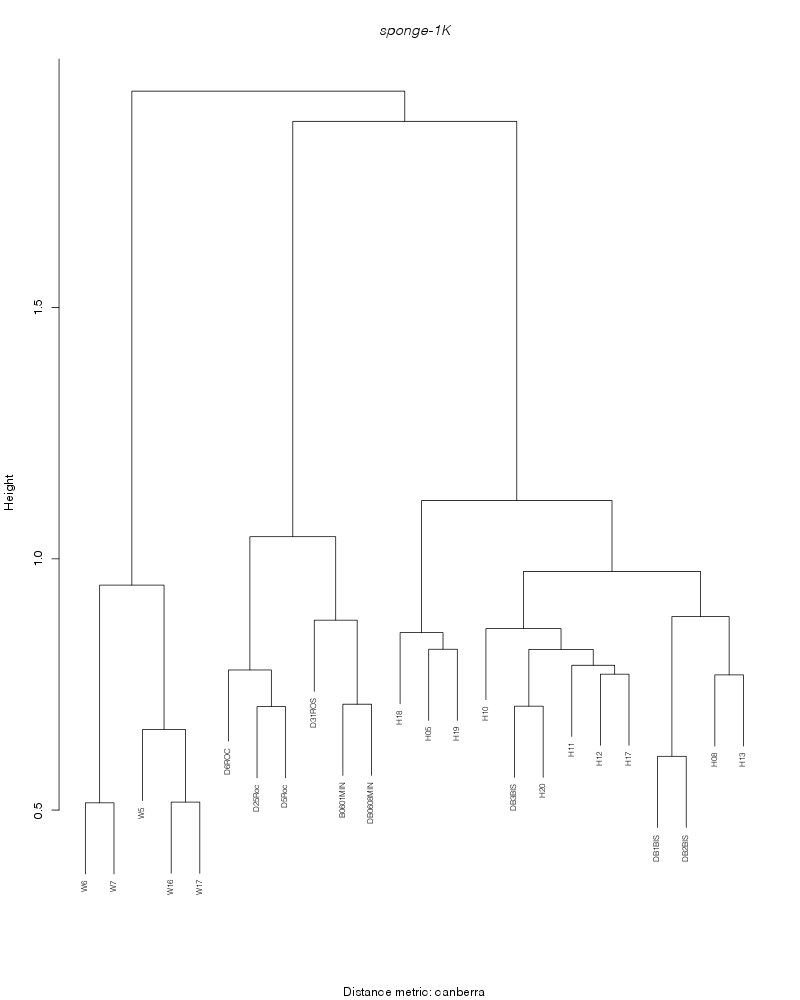
canberra |
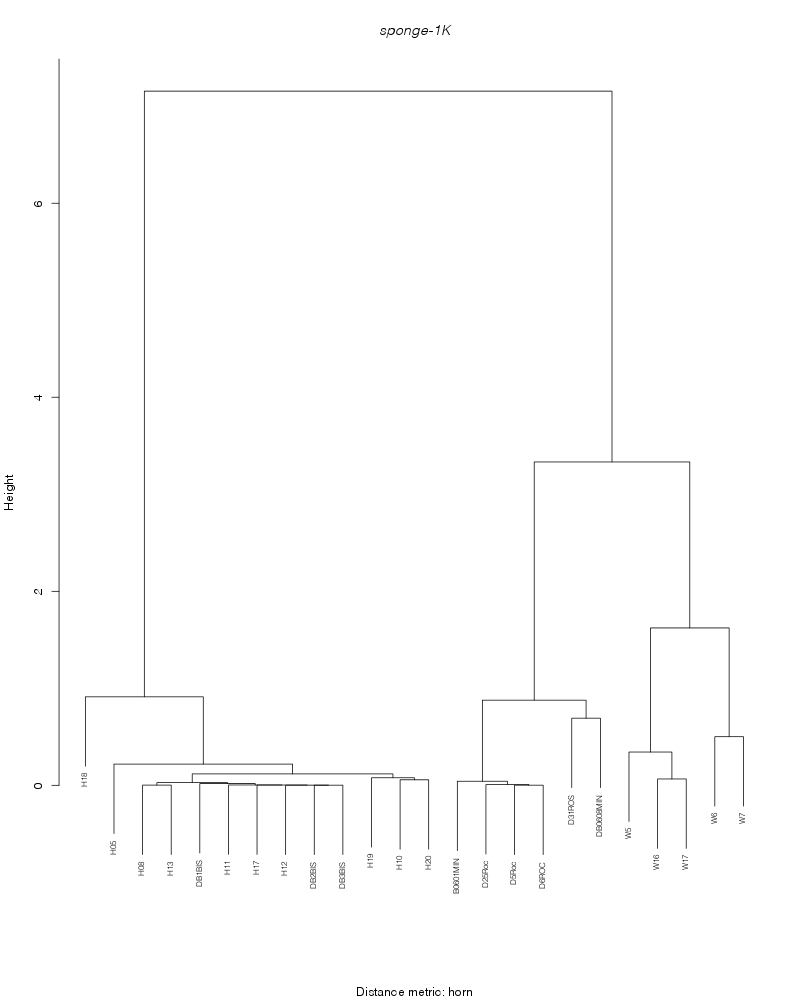
horn |
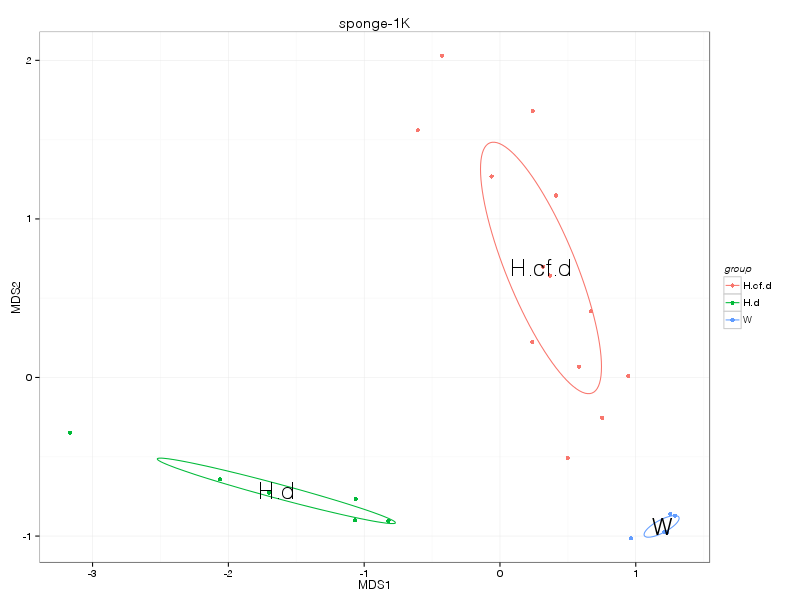
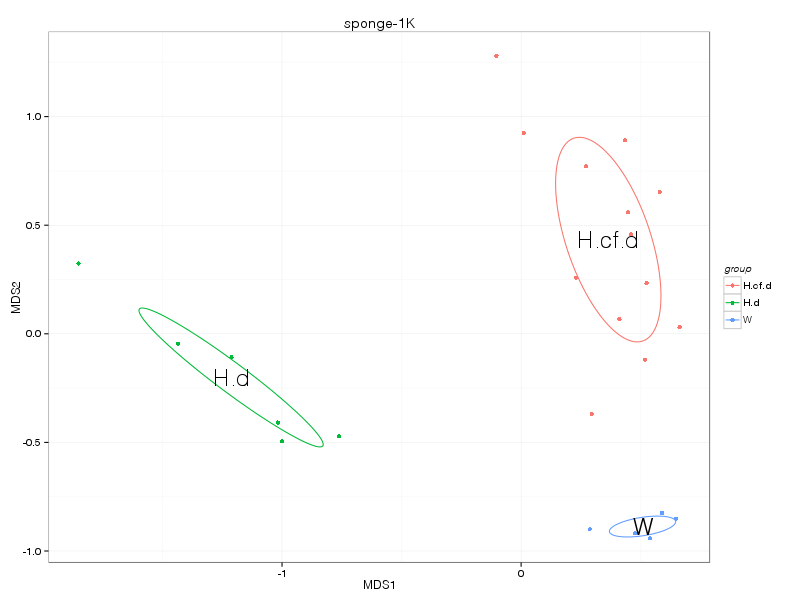
» nmds_analysis
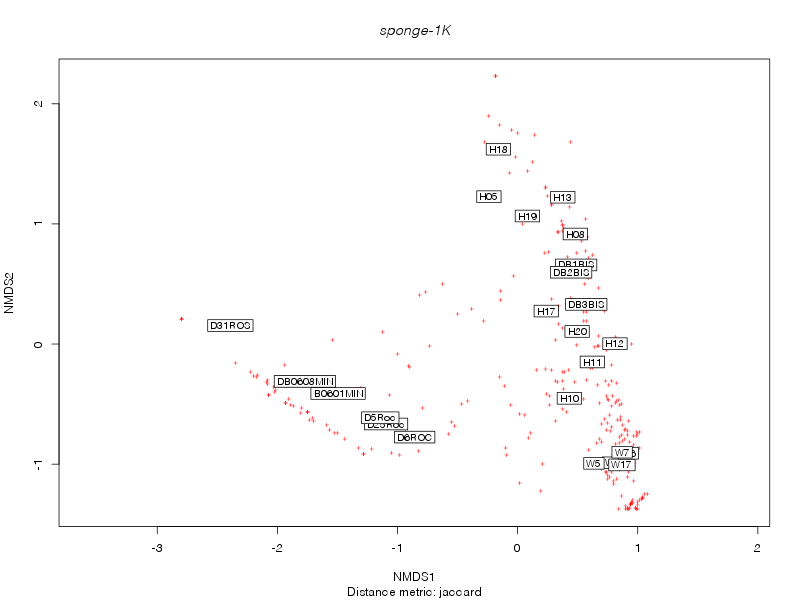
jaccard |
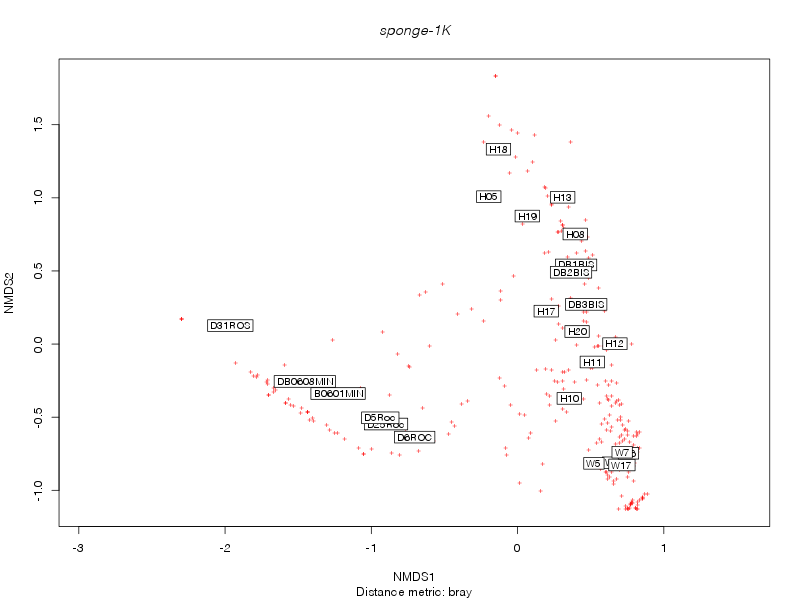
bray |
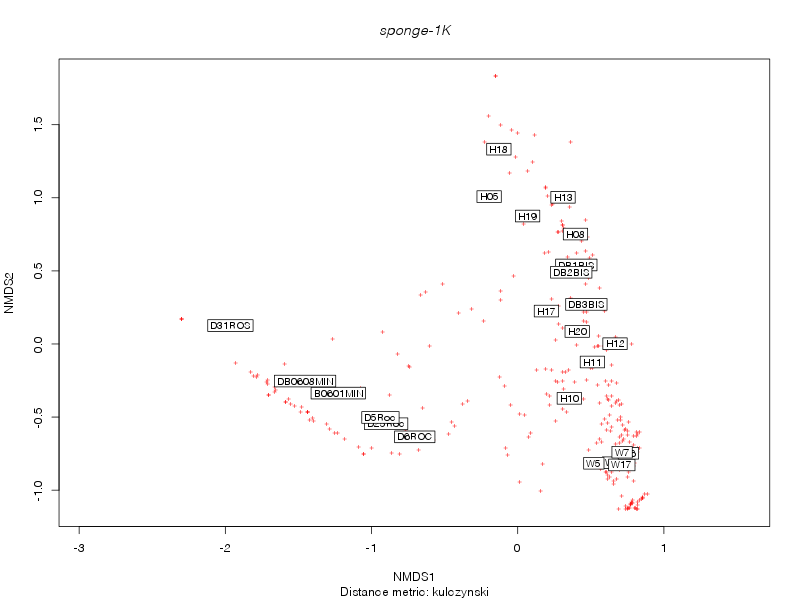
kulczynski |
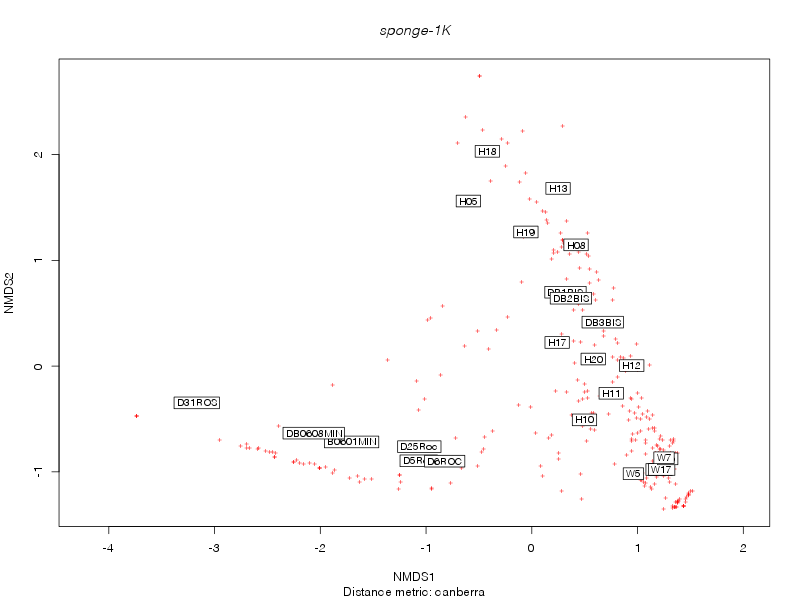
canberra |
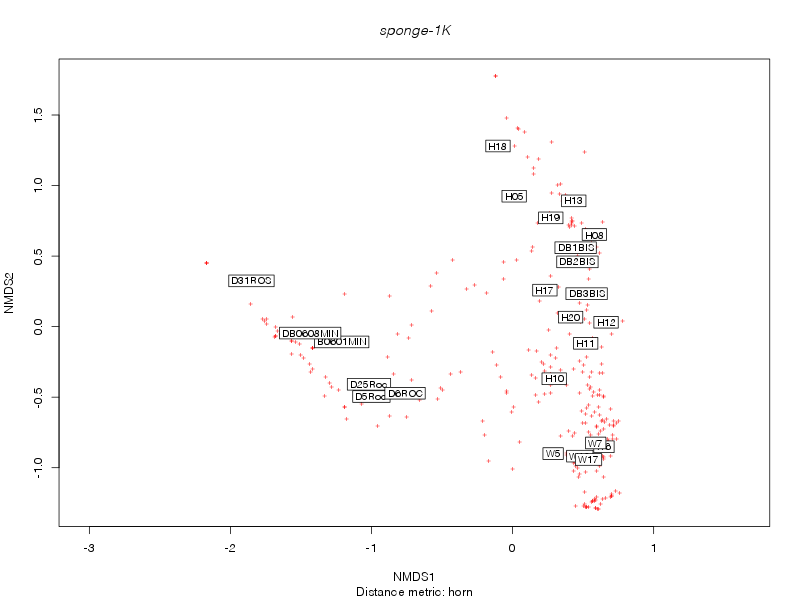
horn |
» SOURCE
» nmds_analysis
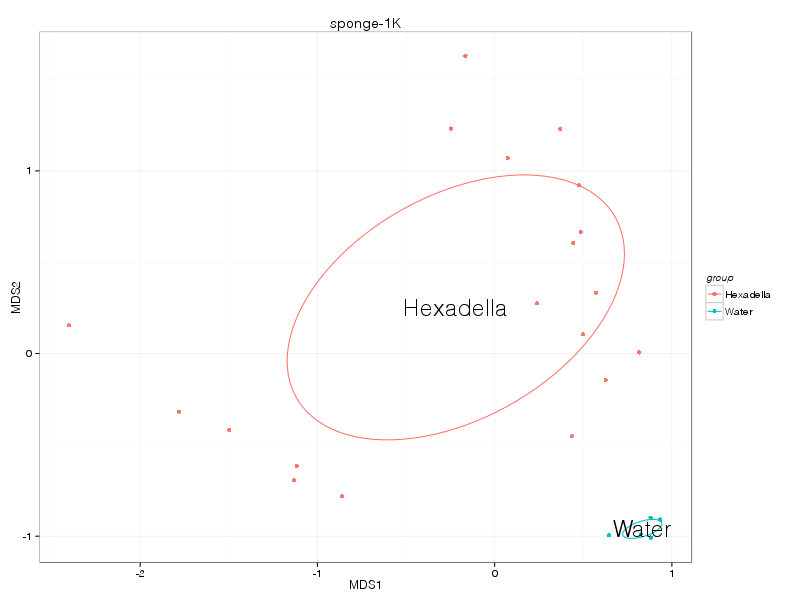
jaccard |
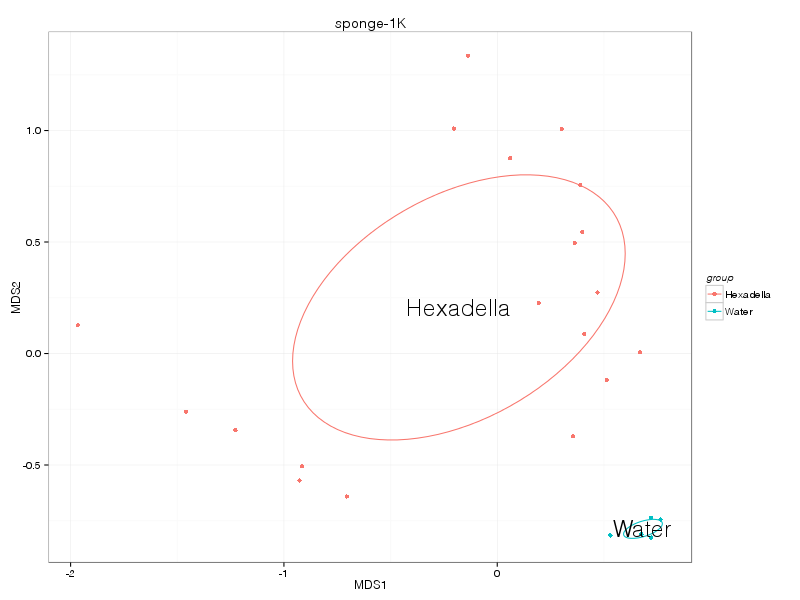
bray |
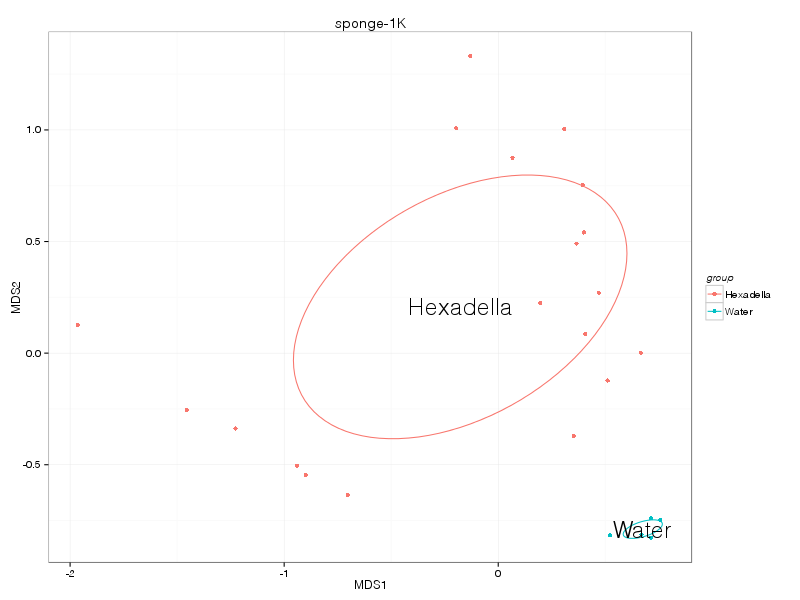
kulczynski |
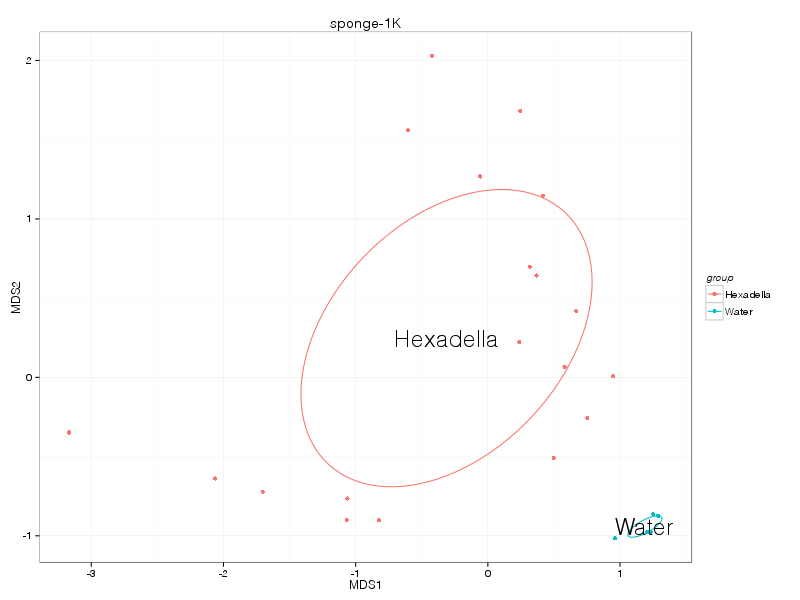
canberra |
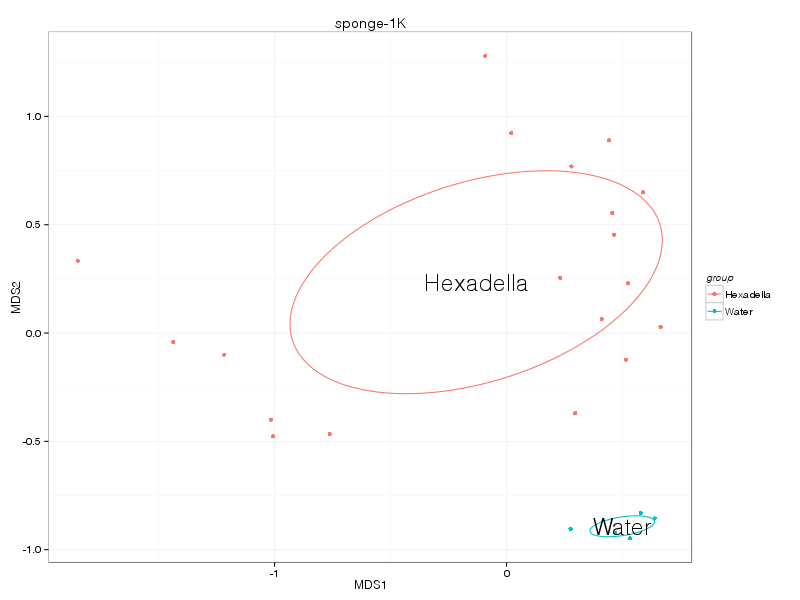
horn |
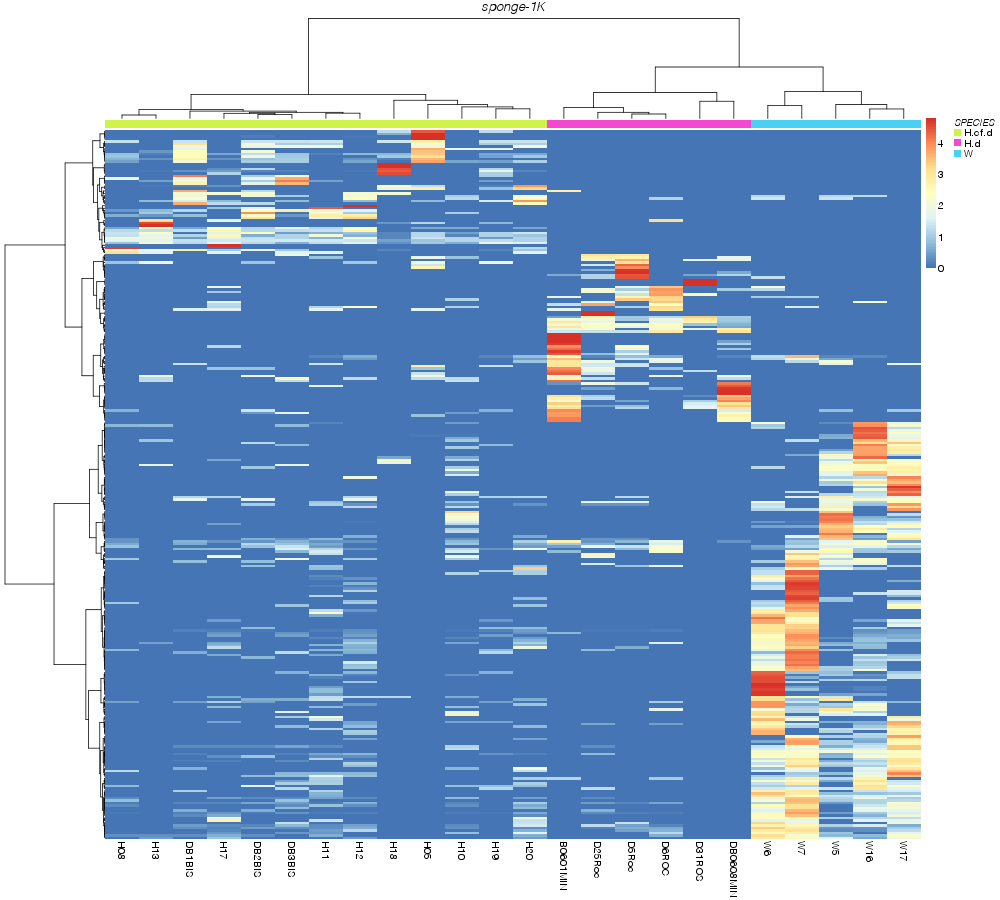
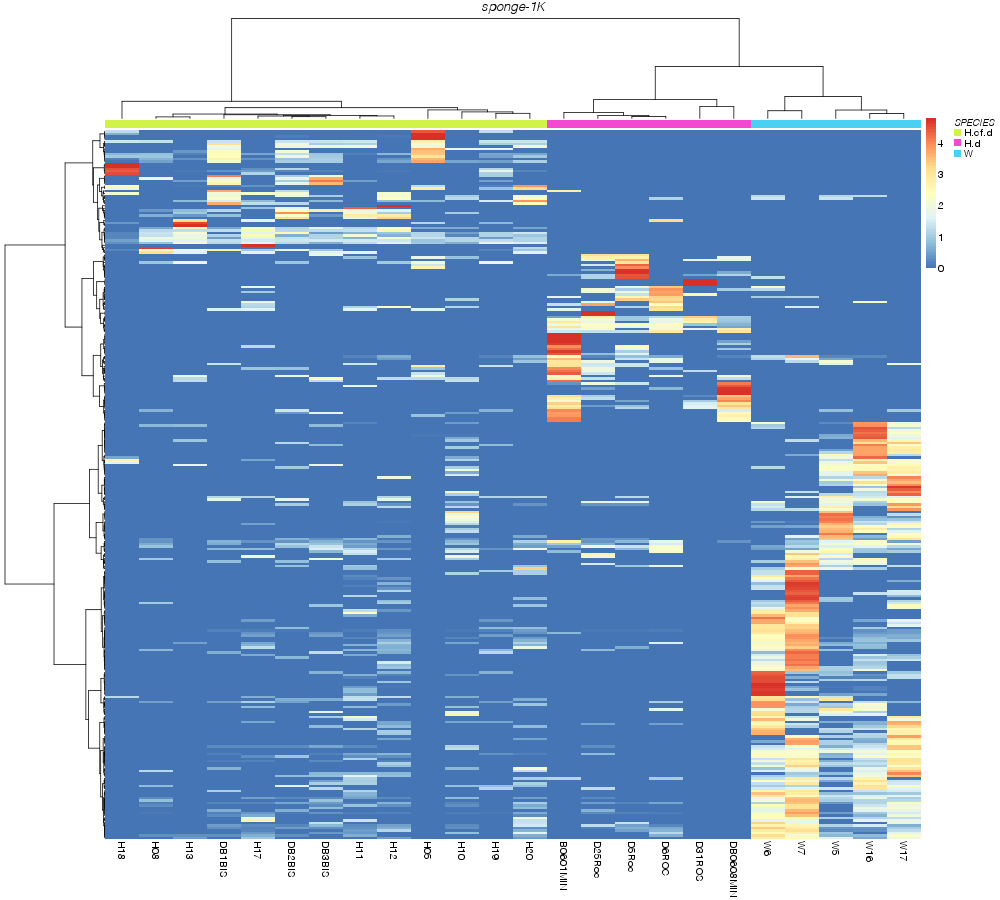
» heatmap_analysis
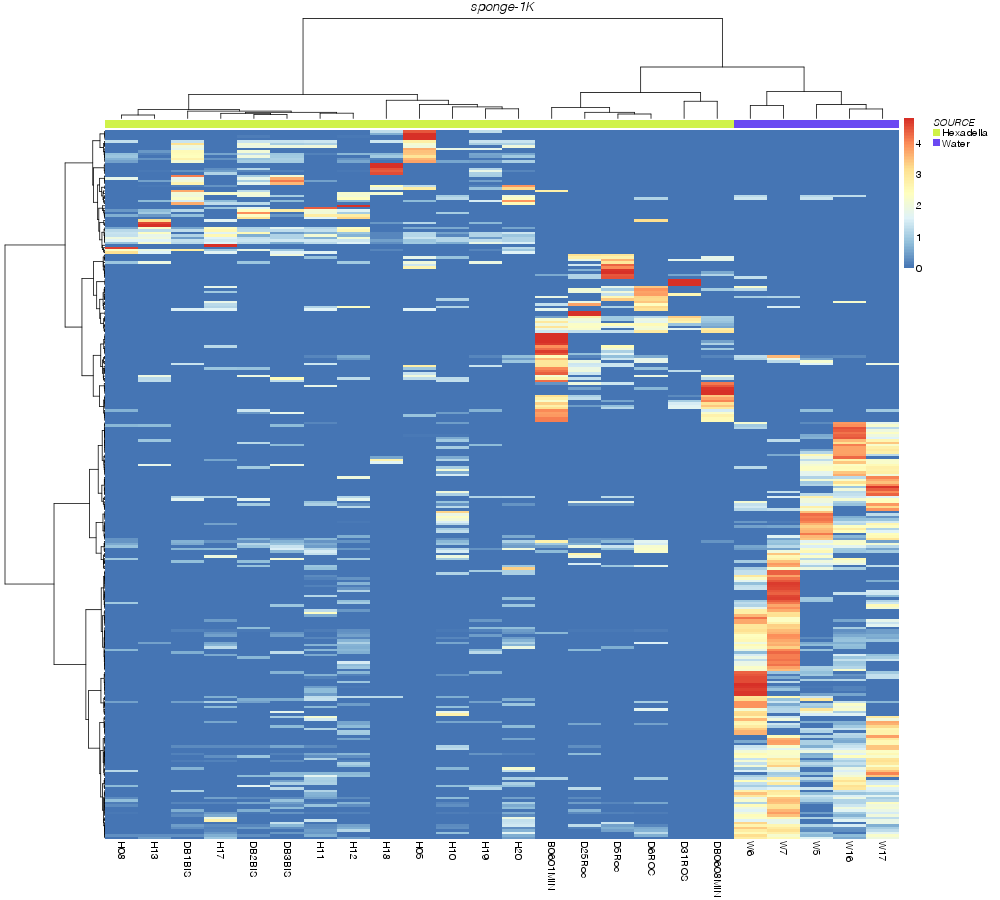
jaccard |
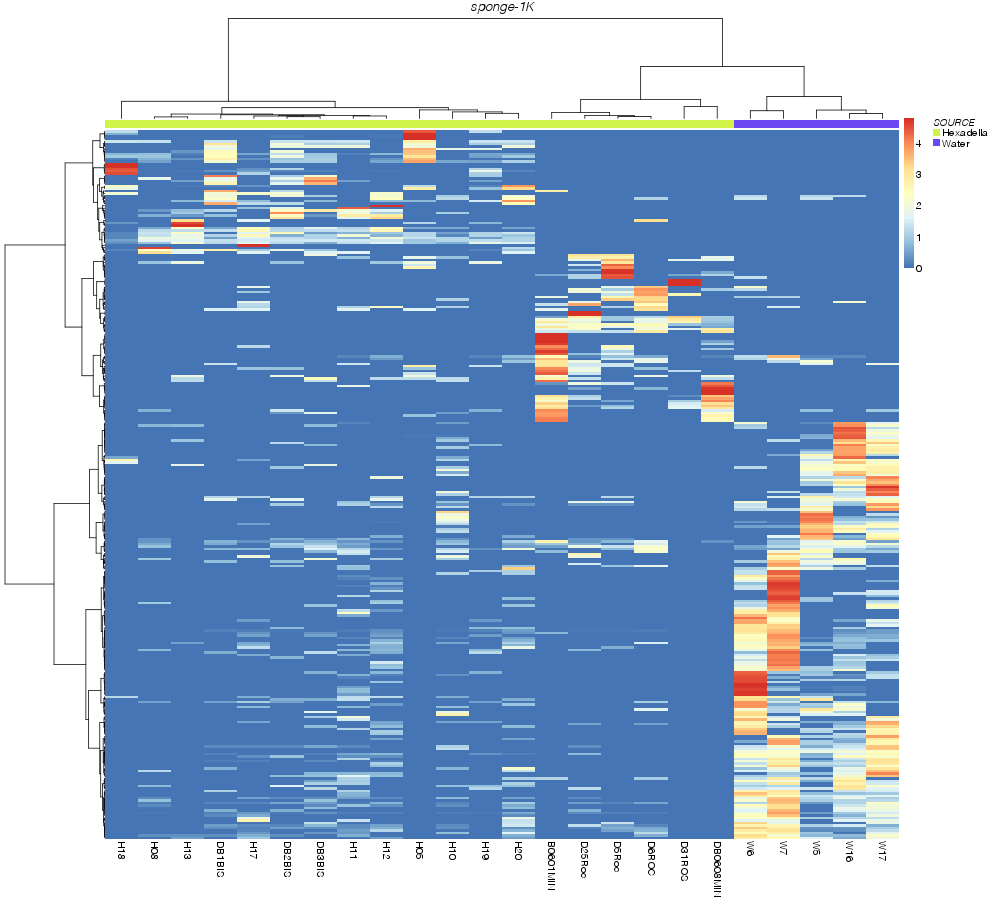
bray |
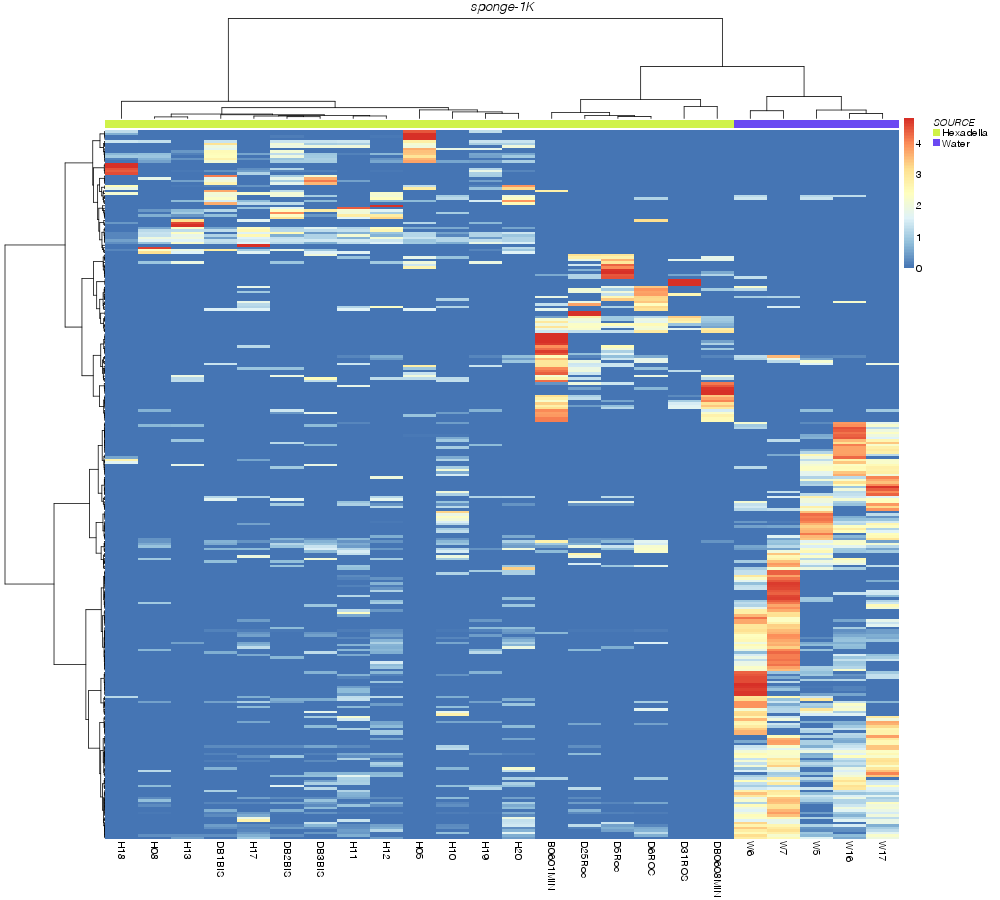
kulczynski |
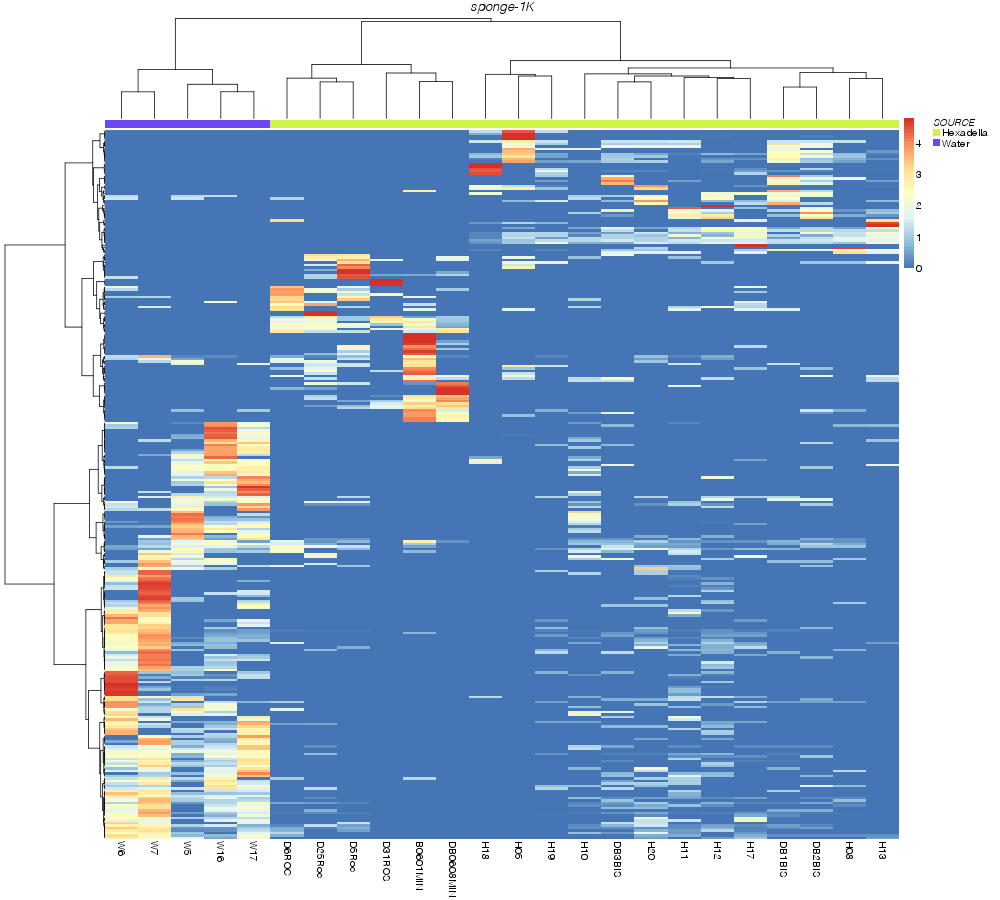
canberra |
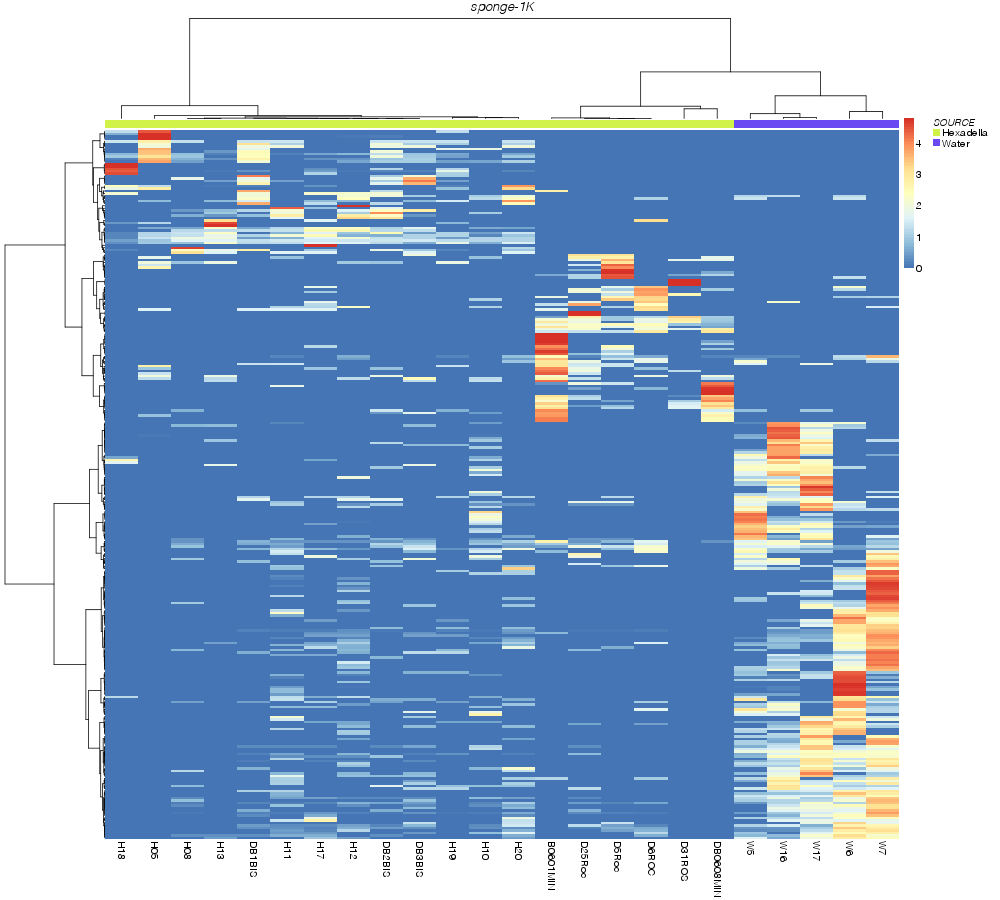
horn |
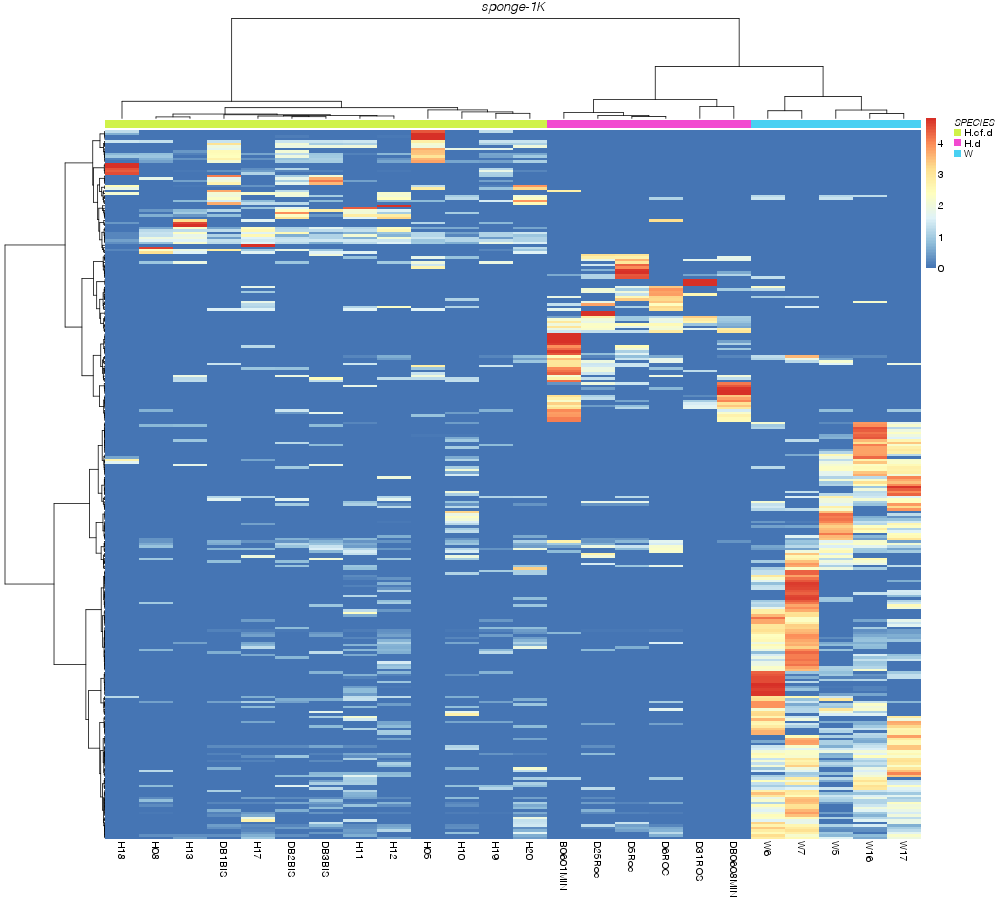
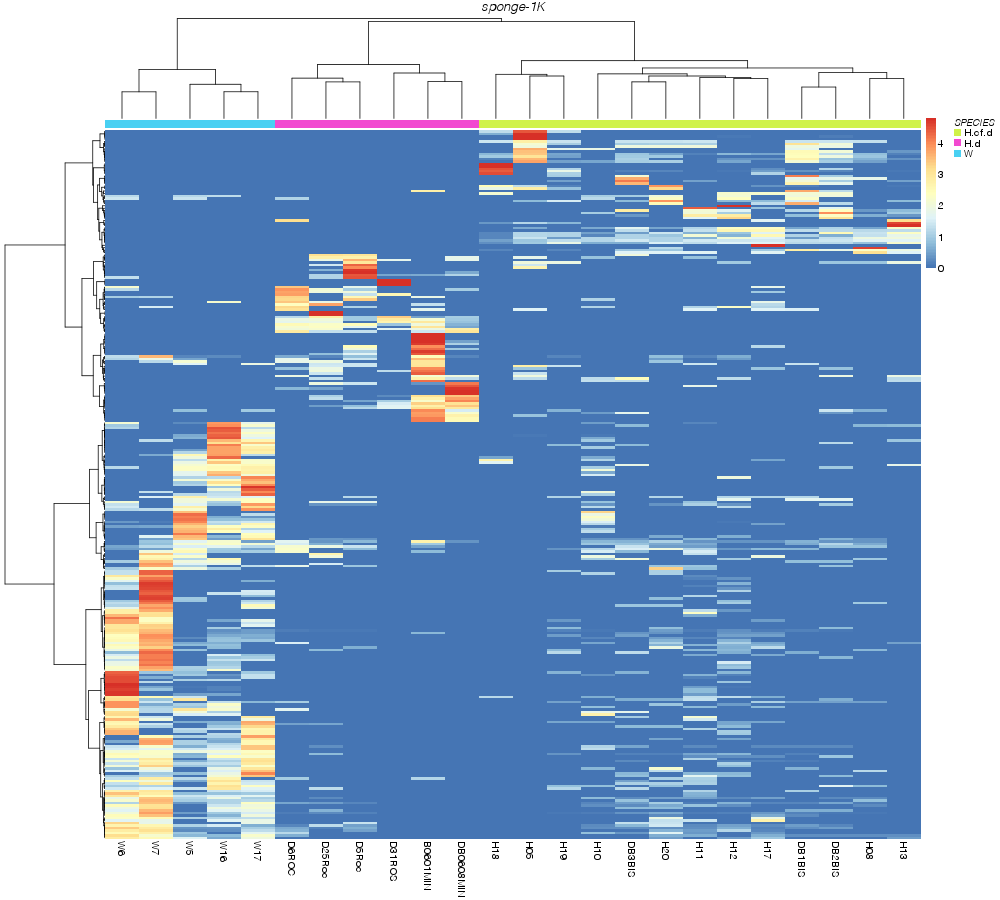
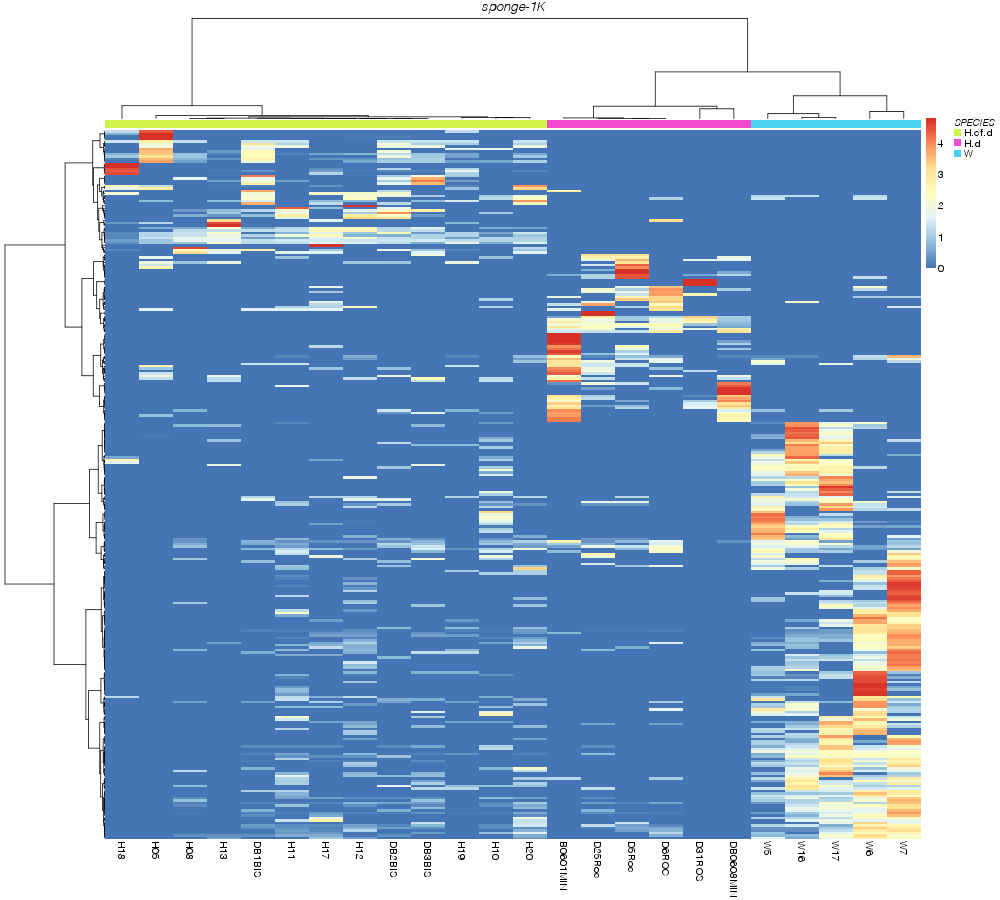
» SPECIES